Oncogenic Rhadinoviruses and Pathogen Discovery
Antiviral Immunotherapy against the human Cytomegalovirus
This research builds on our development of the first CMV-specific CAR (Full et al., 2010; Bednar & Ensser, 2021). We continue this project, which started in a longstanding cooperation with the group of Prof. Wolfgang Holter and PD Dr. Manfred Lehner, St. Anna Children’s Hospital and Children’s Cancer Research Institute, Vienna, within the research training grant GRK2504 project B6. Here we seek proof of concept for antiviral therapies with chimeric antigen receptors in the murine model and mouse CMV (MCMV) (Cordsmeier, 2023b) using novel sensors for T cell mediated cytotoxicity (Bednar, 2023). We are also developing bispecific reagents, based on an earlier cooperation with the biopharmaceutical company Amgen on BiTE®-based immunotherapies for CMV infection (Brey et al., 2018; Menschikowski et al., 2024), for the MCMV model.
Lymphocyte growth transformation by Herpesvirus saimiri (HVS)
Lymphocyte growth transformation by Herpesvirus saimiri (HVS) is a useful tool for immunologists (Bohlen, 2023) as well as a model that allows us to study episomal viral genomes. Our analysis of the viral episomal chromatin structure in human T-cells, together with a seminal characterization of viral episomal DNA replication, can serve to identify factors that regulate chromatin permissiveness within the vector genome and will allow sustained transgene expression from rhadinoviral vectors. We are currently using single cell sequencing to study the viral effectors that HVS uses to transform human and monkey T-cells to antigen independent growth. Primarily we rely on virus-genetic approaches in the context of the viral genome, which we manipulate by standard molecular techniques and homologous recombination. We have shown that constitutive STAT3 activation by the Tip oncogene is not required for human T-cell transformation by HVS and that the Tip-Lck interaction is necessary for transformation and that IL2-dependence of transformation is coupled to a specific tyrosine residue – a finding that directed our research to the T-cellular IL2 signaling pathway and the role of STAT5.
Cellular restriction of the Kaposi sarcoma associated human herpesvirus 8 and related Rhadinoviruses
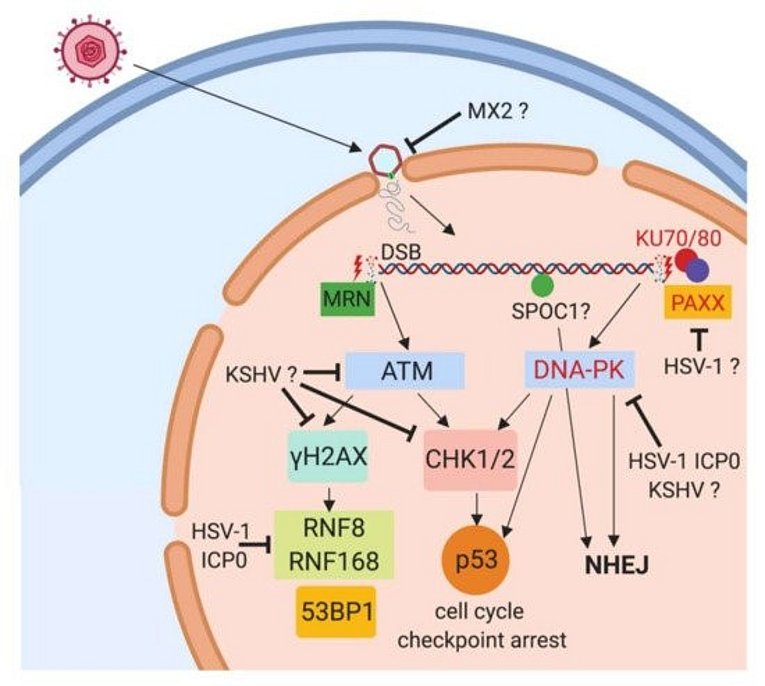
KSHV/HHV8 is a human herpesvirus with close relationship to HVS, Rhesus Rhadinovirus (RRV) is the corresponding homolog from Old World Primates. We are investigating the potential of rhadinoviral proteins to disrupt PML nuclear bodies and other cellular functions such as the DNA Damage Response (DDR), again employing recombinant viruses. This showed that gamma-herpesviral effectors antagonize nuclear domain 10 instituted intrinsic immunity in different ways (Full & Ensser 2019). The HVS ORF3 was able to mediate the selective degradation of the cellular protein Sp100 (Full et al., 2012). The related ORF75 of KSHV was identified as an essential viral protein as it mediates disappearance of ATRX and dispersal of Daxx from ND10 (Full et al., 2014). Notably, the viral ORF75 protein RRV, despite more closely related to KSHV, resembles the HVS ORF3 in its predominant targeting of the major ND10 component SP100 in a proteasome dependent manner (Hahn et al., 2016). In a further search for antiviral restriction factors, our alumni Dr. Florian Full in cooperation with Prof. Michaela Gack demonstrated that the centrosomal protein TRIM43 restricts herpesvirus infection by regulating nuclear lamina integrity (Full et al., 2019).
Pathogen discovery and analysis using next generation sequencing
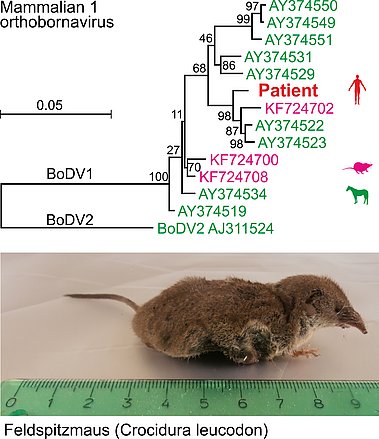
Using next-generation sequencing (NGS), we attempt unbiased pathogen identification from diagnostic samples of patients with presumed diseases of probable infectious origin. Here we detected Borna disease virus 1 (mammalian 1 orthobornavirus, BoDV-1) in brain tissues of patients with fatal encephalitis of previously unknown origin (Korn, 2018; Coras, 2019), demonstrating that BoDV-1 is indeed pathogenic in humans.
We further use NGS for pathogen detection (e.g. Bertz 2021, Stebner, 2021) and in genome wide CRISPR/Cas9 knockout screens (Xiao, 2021), in particular as a platform to search for cellular factors restricting the replication of Herpesviruses and other pathogens.
Recombinant SARS-CoV-2 for vaccine development and analysis of antiviral restriction.
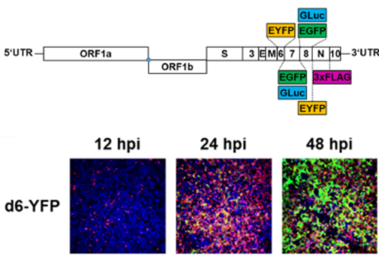
We have cloned and rescued the complete SARS-CoV-2 virus genome in a bacterial artificial chromosome (BAC) vector, directly from diagnostic material without prior cell culture passage (Hermann 2021; Cordsmeier 2023a). Using derived marker viruses, we analyze innate and cellular restriction of SARS-CoV-2, with a focus on TRIM family proteins. Recombinant viruses are successfully used in cooperative projects (e.g. Chen 2023; Kim 2023; Müller 2023; Wu 2022; Zech 2021). Furthermore, viruses are attenuated in their suppression of innate and cellular responses by introduction of specific mutants. These are investigated regarding their suitability as prototypic live attenuated viruses (LAV).