The viral genome associated proteome (VGAP)
Dr. Florian Full, Marie Skłodowska-Curie Individual Fellowship
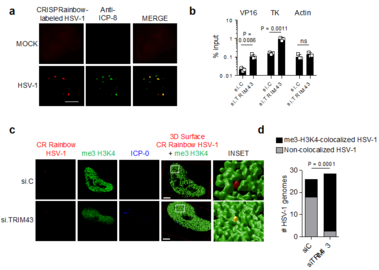
a) Co-localization of HSV-1 genomes with the viral DNA-binding protein ICP8. HeLa cells expressing the CRISPRainbow (CRainbow) components dCAS9, a HSV-1 genome-specific sgRNA and a sgRNA-binding blue-fluorescent protein (shown in red), were infected with HSV-1 (MOI 1) for 1.5 h, followed by immunostaining with antibodies specific for ICP8 (green). b) Chromatin-IP (ChIP) analysis of HFF cells transfected with the indicated siRNAs for 48 h and then infected with HSV-1 (MOI 1) for 5 h using an anti-me3-H3K4 antibody. ChIP DNA was quantified by qPCR with promoter-specific primers to viral genes VP16 and TK as well as cellular actin (control). Data were normalized to the level of input DNA and presented as percentage of input. c) Co-localization of HSV-1 genomes with me3-H3K4 chromatin using CRainbow. HeLa cells expressing the CRainbow components dCAS9, a HSV-1 genome-specific sgRNA and a sgRNA-binding blue-fluorescent protein (shown in red), were transfected with si.C or si.TRIM43. 24 h later, cells were infected with HSV-1 (MOI 1) for 1.5 h, followed by immunostaining with antibodies specific for me3-H3K4 (green) and ICP-0 (blue). Stacks of confocal images were subjected to 3D surface rendering. Scale bar, 3 µm. d) Quantification of the co-localization of HSV-1 genomes with me3-H3K4 for the experiment shown in (c). n = 25 cells.
Data represent mean and s.d. of n = 3 (b), and statistical significance was calculated using unpaired two-tailed students t-test (b) or two-way ANOVA test (d).
(from Full et al., Nature Microbiology, 2019).
Herpesviruses have evolved sophisticated ways to subvert the immune system during millions of years of coevolution with their respective hosts. For Herpesviruses the antagonism of the innate immune system is of particular importance in order to achieve an efficient infection of target cells. The nine human members of the herpesvirus family include important human pathogens like Herpes-simplex viruses-1 and -2 (HSV-1 and HSV-2), Varicella-Zoster Virus (VZV), Human Cytomegalovirus (HCMV), Human Herpesviruses-6A, -6B and -7 and the tumor viruses Epstein-Barr virus (EBV) and Kaposi's sarcoma associated herpesvirus (KSHV). A hallmark of herpesvirus infections is the establishment of lifelong latency, for example about 3.7 billion people alone are infected with HSV-1 worldwide, and even more by HHV-6, -7 and EBV. In this regard herpesviruses are the most "successful" virus family in the human population, because presumably almost all human adults are latently infected with at least one herpesvirus and most of us by several. Herpesviruses are double-stranded DNA viruses with genome sizes from 125-250 kbp. For infection of target cells, herpesviral virus particles bind to cellular surface receptors. Upon infection the viral capsid is transported through the cytoplasm along the cytoskeleton until it reaches the nuclear membrane; there the viral genome, which is coiled in the capsid under high pressure, is injected through nuclear pores. Whereas the viral genome is enclosed and protected by capsid proteins in the cytoplasm, upon injection into the nucleus the naked linear viral DNA is especially vulnerable for detection by cellular DNA-damage proteins and attack by cellular restriction factors. Therefore the viruses had to evolve ways to i) prevent detection of the genome by DNA-damage proteins; ii) prevent degradation of the naked, linear viral DNA; iii) realize recircularization of the viral genome; iv) achieve chromatinization of the genome but simultaneously prevent the addition of repressive chromatin marks and vi) establish efficient viral gene expression.
The goal of this project was the identification of novel restriction or host factors of herpesviruses that are involved or interfere with the processes mentioned above. Therefore a new method called HyCCAPP should be developed in order to discover new factors that associate with the viral genome upon nuclear entry. Although the initially proposed approach failed, we used alternative methods, namely CRISPRainbow- and EdC-labelling of viral genomes, that allowed us to identify SMCHD1 as a novel restriction factor for herpesviruses that is associated with viral genomes. SMCHD1 is an important human protein, and mutations in SMCHD1 have been shown to be the cause of two genetic diseases, Facioscapulohumeral muscular dystrophy (FSHD) and Bosma arhinia microphtalmia syndrome (BAMS). We characterized SMCHD1 in detail as antiviral protein and could even show viral antagonism of SMCHD1-mediated restriction by a protein of KSHV, further corroborating the importance of SMCHD1 in antiviral immunity. We hope that our discovery of a new antiviral restriction factor may help to understand the pathogenicity of herpesviral disease and contribute to the development of new anti-herpesviral drugs in the future.
Full, F., M. van Gent, K. M. J. Sparrer, C. Chiang, M. A. Zurenski, M. Scherer, N. H. Brockmeyer, L. Heinzerling, M. Sturzl, K. Korn, T. Stamminger, A. Ensser, and M. U. Gack. "Centrosomal Protein Trim43 Restricts Herpesvirus Infection by Regulating Nuclear Lamina Integrity." Nat Microbiol 4, no. 1 (2019): 164-76.
Florian Full (MSC Fellow), Armin Ensser

This project was funded by the European Union
Marie Skłodowska-Curie Individual Fellowship