Protective Immune Response against Cytomegalovirus
Human cytomegalovirus (HCMV) is an important, ubiquitously occurring, human pathogen in immunocompromized hosts. The virus can cause severe disease in transplant recipients. In large parts of the world, HCMV is also the most common viral infection acquired in utero. In the USA and Europe an estimated 0.2% -1.2% of all live born infants are infected with HCMV and 5-15% of these babies develop long term sequelae resulting from congenital infection. As a consequence of the importance of congenital HCMV infection for public health, the Institute of Medicine at the National Institutes of Health (NIH), USA, has ranked the development of an HCMV vaccine as a top priority. A major obstacle for the development of a vaccine is a lack of knowledge of the nature and specificities of protective responses that should be induced by the vaccine. HCMV is a complex virus containing numerous antigens within the viral envelope that could be targets for protective antibodies. The fusogen glycoprotein B (gB) and/or the heterodimeric gH/gL which are components of the receptor binding trimeric (gH/gL/gO) or pentameric (gH/gL/UL128,130,131A) complexes are important targets for neutralizing antibodies and hence interesting molecules for intervention strategies such as vaccination or passive immunotherapy.
The main research areas of our lab are:
- Uncovering novel neutralizing epitopes on cytomegaloviral glycoproteins
- Define the capacity of antibodies to limit cell-associated spread of HCMV
- Obtain molecular insights into the fusion process of HCMV-gB or SARS-CoV-2 spike protein
- Identification of antibody signatures as predictors for HCMV disease and transmission
- Characterization of the mechanisms of protection in vivo
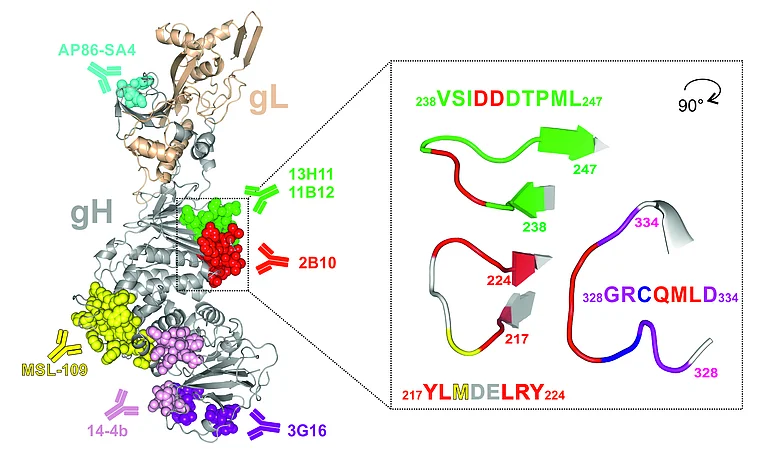
Known binding sites of gH-specific antibodies isolated from mouse (AP86-SA4, 14-4b, as well as 6E3, 3C11, and 2B10), and human (13H11, 11B12, MSL-109, and 3G16) are indicated with their respective epitopes within the ribbon model of gH (gray) complexed with gL (wheat) as deduced from Chandramouli et al. 2017 (PDB ID 5VOB). Right: Detailed presentation of the 2B10 epitope, rotated 90 degrees around the vertical axis relative to its orientation in the left panel. Amino acids crucial for 2B10 binding are highlighted in red, and Met in yellow. (Modified from Thomas M., et al., J Virol. 2021).
Uncovering novel neutralizing epitopes on cytomegaloviral glycoproteins
To extend the knowledge of mechanisms of virus neutralization, monoclonal antibodies (mAbs) were generated following immunization of mice with HCMV. Hybridoma supernatants were screened for in vitro neutralization activity, yielding three potent mAbs, 6E3, 3C11, and 2B10 (Figure 1). MAbs 6E3 and 3C11 blocked infection of all viral strains that were tested, while mAb 2B10 neutralized only 50% of the HCMV strains analyzed. The strain-specific determinant for neutralization by mAb 2B10 was identified as a single Met/Ile amino acid polymorphism within gH, located within the central part of the protein. The polymorphism is evenly distributed among described HCMV strains.
Thus, the 2B10 epitope represents a novel strain-specific antibody target site on gH of HCMV. The dependence of the reactivity of 2B10 on the simultaneous presence of gB/gH/gL will be of value in the structural definition of this tripartite complex. The 2B10 epitope may also represent a valuable tool for diagnostics to monitor infections/reinfections with different HCMV strains during pregnancy or after transplantation.
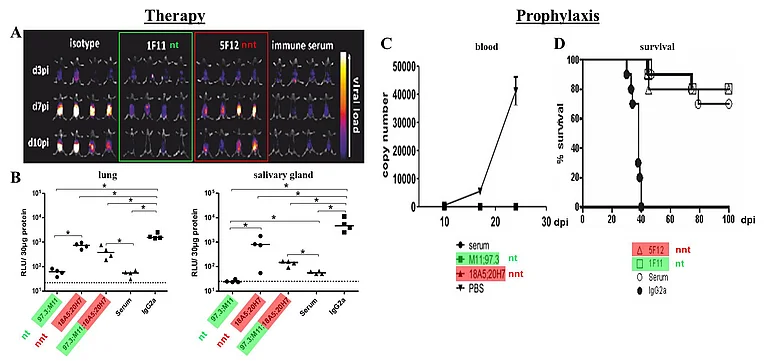
(A+B) Viral load of RAG-/- mice after therapy with nt or nnt antibodies or combinations of both. Mice were infected with 105 pfu MCMV157luc and treated with a total of 250 μg of the indicated IgG(s) or 200 μl serum per animal at three days after infection. (A) In vivo imaging was performed at the indicated days post infection (dpi). (B) Viral load in aliquots of organ homogenates containing 30 μg protein was determined 10 days after infection by a luciferase based assay. RLU: relative light units. Statistics: One way ANOVA using Bonferroni´s multiple comparison test *: p<0.05., dotted line: detection limit. (C+D) Protective capacity of anti-gB mabs following prophylactic application. A total of 250 μg IgG(s) per mouse or 200 μl serum/PBS was injected one day before infection with 104 pfu of MCMV157luc. (C) Blood was taken at the indicated dpi and MCMV load determined by qPCR. n = 4 in antibody treated groups and n = 3 in the PBS treated group. Values represent mean (SEM) of all mice within one group and duplicate determinations per sample. MCMV genome copy number is given per 1 μg total DNA. Detection limit: 1 copy/50 ng total DNA. (D) Survival was monitored for 100 days p.i. Statistics: log-rank (Mantel-Cox) test: p <0.0001. Representative data from 2 independent experiments.
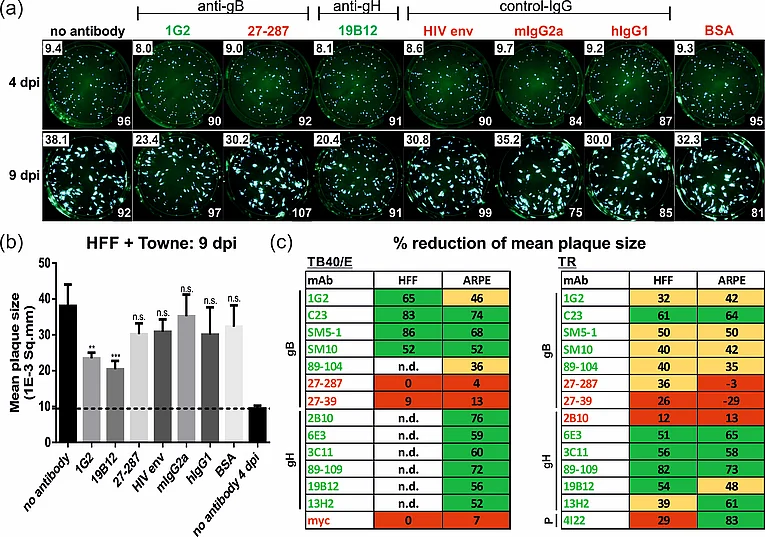
HFFs were infected with HCMV strain Towne. After 24 h, medium was replaced by agarose overlay medium containing the antibodies (50 µg/mL) or BSA, respectively. Starting at 4 dpi, images were captured by a Fluorospot reader and used for automated quantification of the mean plaque size (1E-3 Sq.mm, numbers in the upper left corner) of all fluorescent spots (numbers in the lower right corner). (b) Bar graphs of the mean plaque sizes at 9 dpi. (c) Capacity of the indicated mAbs to reduce the mean plaque size (in % relative to the no antibody control at 4 dpi) following infection of HFF or ARPE cells with strains TB40/E or TR. Red, not significant (<30%); yellow, significant (30–49%); green, highly significant reduction (>50%); n.d., not determined. (Modified from Reuter et. al., Viruses, 2022.)
Define the capacity of antibodies to limit cell-associated spread of HCMV
In vivo HCMV spreads cell-to-cell. Thus, quantifying the inhibition of cell-to-cell transmission could be important in the evaluation of therapeutic antibodies and/or humoral responses to infection or immunization. We established a quantitative plaque reduction assay, which allowed for the measurement of the capacity of antibodies to limit HCMV spread in vitro. Using an automated Fluorospot reader, infection progression was assayed by the expansion of viral plaques during the course of infection with various GFP-expressing viruses. We found that in contrast to non-neutralizing mAbs, neutralizing mAbs against both glycoprotein B and H (gB and gH) could significantly inhibit viral plaque expansion of different HCMV strains and was equally efficient in fibroblasts as in epithelial cells (Figure 2). In contrast, an anti-pentamer mAb was active only in epithelial cells. Taken together, our data demonstrate that specific anti-HCMV mAbs can significantly limit cell-associated virus spread in vitro. A better understanding of the target epitopes targeted by antibodies that block cell-associated HCMV transmission will likely be of great importance in the design of improved vaccine or therapeutic antibodies for prophylaxis and treatment of HCMV infections.
Obtain molecular insights into the fusion process of HCMV-gB or SARS-CoV-2 spike protein
One main interest of our lab is to define the mechanisms of protective antibodies against the fusion protein gB of HCMV. By utilizing a panel of virus-neutralizing gB-specific monoclonal antibodies (MAbs), it was demonstrated that syncytium formation of an intrinsically fusion-active gB/VSV-G chimera was inhibited by only a subset of neutralizing MAbs, which target a distinct antigenic domain of gB. This observation argues for differential modes of action of HCMV neutralizing anti-gB MAbs and suggests that blocking the membrane fusion function of gB could be one mechanism of antibody-mediated virus neutralization.
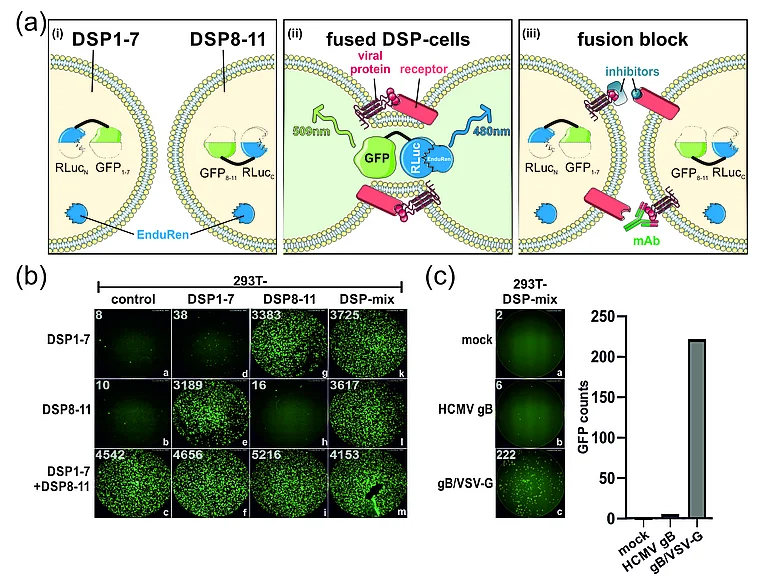
Like HCMV, the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) spreads efficiently cell-to-cell through mechanisms facilitated by its glycoprotein spike. In order to dissect the cell membrane fusion activity of spike in further detail, we established a dual split protein (DSP) assay based on complementation of GFP and luciferase, to quantify the fusogenic activity of the viral fusion proteins (Figure 3). We provide several lines of evidence that the spike protein of SARS-CoV-2, but not SARS-CoV-1, induced cell–cell fusion even in the absence of its receptor, angiotensin-converting enzyme 2 (ACE2).This poorly described ACE2-independent cell fusion activity of the spike protein was strictly dependent on the proteasomal cleavage of spike by furin, while TMPRSS2 was dispensable. Previous and current variants of concern (VOCs) differed significantly in their fusogenicity. The Delta spike was extremely potent compared to Alpha, Beta, Gamma and Kappa, while Omicron spike was almost devoid of receptor-independent fusion activity. Nonetheless, for all analyzed variants, cell fusion was dependent on furin cleavage and could be pharmacologically inhibited with CMK. Mapping studies revealed that amino acids 652-1273 conferred the ACE2-independent fusion activity of spike. Unexpectedly, residues proximal to the furin cleavage site were not of major relevance, whereas residue 655 critically regulated fusion. Finally, we found that spike’s fusion activity in the absence of ACE2 could be inhibited by antibodies directed against its N-terminal domain (NTD) but not by antibodies targeting its receptor-binding domain (RBD). The DSP assay allows screening for inhibitors or antibodies that interfere with the fusion process and may therefore contribute to both rational vaccine design and development of novel treatment options against the human pathogenic viruses SARS-CoV-2 or HCMV.
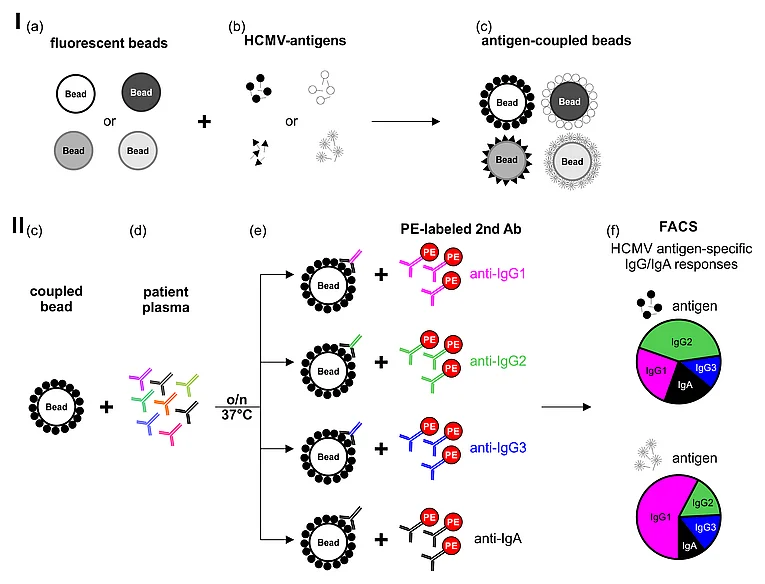
(I) (a) paramagnetic fluorescent beads loaded with E-coil peptides are coupled via Bind & Bite technology with (b) HCMV antigens fused with a C-terminal K-coil-tag. (II) (c) Antigen-coupled beads are (d) incubated with human sera followed by (e) incubation with PE-coupled secondary antibodies. (f) The antigen-specific antibody responses are determined by FACS and/or Luminex assay.
Identification of antibody signatures as predictors for HCMV disease and transmission
HCMV primary infection, reactivation, or reinfection can hardly be discriminated and predictions for the risk of HCMV transmission and likelihood to progress to disease cannot be made. Our aim is to identify HCMV antibody signatures induced by various HCMV antigens that might serve as predictors for HCMV disease and transmission. As an alternative to inactivated virus particles, recombinant viral proteins are increasingly being used as antigens for serological test methods. However, their coupling to surfaces require complex development and validation work. A variable that is difficult to control is the spatial orientation and presentation of the proteins immobilized on the surfaces used. Specific hetero-dimerizing peptide tags, so-called coiled-coils (Bind & Bite technology), offer a clear improvement.
For this, heterodimers of two peptides with K-coil or E-coil are formed and can be stabilized by covalent coupling. Various structural and non-structural HCMV antigens were cloned for prokaryotic or eukaryotic expression in-frame to a C-terminal K-coil-tag. Via cooperation with our industry partner Virion\Serion (Würzburg, Germany), we were able to purify recombinant HCMV proteins via Bind & Bite technology with E-coil-coupled paramagnetic beads from crude cell lysates (Figure 4). As a proof-of-principle, antigen-loaded magnetic beads were incubated with HCMV-positive sera and stained with a PE-labeled secondary antibody to determine the IgG/IgM-specific antibody responses. As these pilot experiments successfully reconfirmed the seropositive individuals, the set of analyzed HCMV antigens is currently extended, as it will be for the IgG-subclass-specific antibody responses and FcR-binding capacity of the respective sera.
Characterization of the mechanisms of protection in vivo
Using murine cytomegalovirus (MCMV) as a model system, we could demonstrate that adoptive transfer of antibodies can protect immunodeficient RAG-/- mice from the lethal course of MCMV-infection. Strikingly. in vivo protection was not directly correlated to the transfer of virus-neutralizing activity, since (i) mAbs without neutralizing activity in vitro can provide protection in vivo (Bootz et al., 2017), and (ii) Fcγ-receptor-deficient RAG-/- mice could not be protected by antibodies, indicating a decisive role for Fc-mediated effector functions for the protection. To characterize the protective mechanism of mAbs, we have generated an extensive set of neutralizing (nt) and non-neutralizing (nnt) mAbs expressed as different IgG subtypes with or without Fc-mutations that enhance or diminish the capacity of binding to Fcγ-receptors or complement. The potency of these mAbs is currently analyzed by monitoring infection in immunodeficient RAG-/- mice infected with a luciferase- expressing MCMV-strain. Taken together this project assesses the mechanisms of protection facilitated by antibodies as the basis for a rational vaccine design against HCMV.