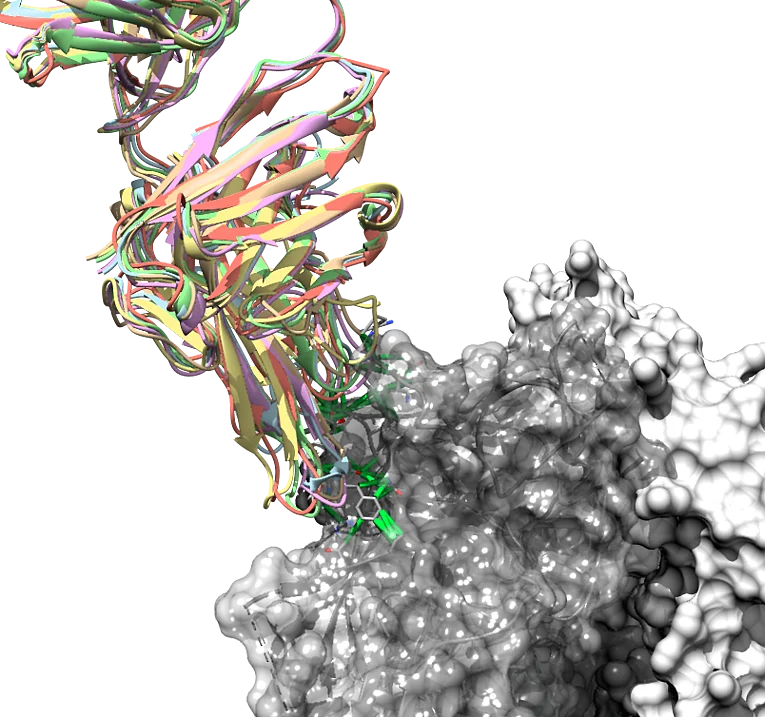
Structure-based design and optimization of ligands for novel antiviral strategies
Antibodies are the multitool of the adaptive immune system. These proteins can identify pathogens and small molecules and either disable them or mark them for elimination by other parts of the immune system. Even though our genes only carry templates for a small number of antibodies the body can modify these templates in a process called affinity maturation. This process allows the generation of antibodies that can identify most harmful substances to the body.
This project focuses on antibodies recognizing the human immunodeficiency virus (HIV) that leads to the acquired immunodeficiency syndrome (AIDS). These antibodies identify HIV by binding to the so-called envelope protein on the surface of HIV.
Unfortunately, HIV is known to mutate frequently and change the appearance of the envelope protein which reduces the efficiency of previously formed antibodies. This makes finding a cure or a vaccine for HIV challenging. However, there are some antibodies that are still able to identify and inactivate the envelope protein even after mutation. These kinds of antibodies are called broadly neutralizing antibodies (bnAbs) and are particularly interesting for researchers because they can serve as a template for new drugs and aid in vaccine design.
We want to investigate the structure of these antibodies and use their key features for the design of new peptide drugs. This approach is cheaper and more flexible than approaches using antibodies because antibodies are generally expensive and time-consuming in large-scale production. To achieve this goal, known structures of anti-HIV antibodies in complex with the envelope protein will be retrieved from protein structure databases such as the Protein Data Bank. The antibodies in these structures will then be evaluated for their binding properties to the viral envelope protein. Candidates that appear suitable for peptide design will be further investigated using molecular dynamics (MD) simulations. MD simulations are a computational method that enables the calculation of movements and interactions of (bio-)molecules. With this, the stability of the interactions between the antibody and the envelope protein can be visualized and analyzed further. The insights gained from these analyses will guide the design of peptides that can replicate the properties of bnAbs, their natural templates, and additionally may help to understand the changes during affinity maturation.