Long and super-long CDR3 regions in human and animal antibodies
Our aim is to understand the selection of motifs in long CDR3 created during somatic hypermutation by quantifying the differences between naïve and memory heavy chain B cell repertoires obtained from next generation sequencing (NGS) and evaluate these differences in model antibodies with computational methods developed for protein structure analysis. In cases where there is no software solution, we aim to adapt existing software or develop novel software solutions.
Next generation sequencing analysis of distinctive B cell subsets from leukapheresis products of hematopoietic stem cells from healthy donors proved to be a powerful tool to characterize the human antibody repertoire. The antibodies produced by human naïve and memory B cells show significant differences in their heavy chain CDR3 repertoires.
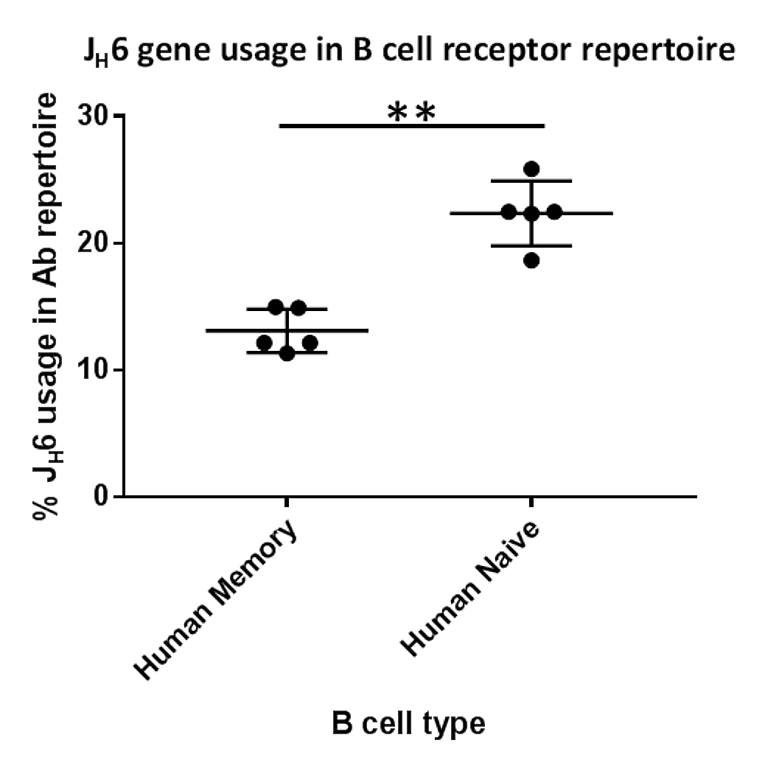
We prove that human heavy chain CDR3 loops are shorter in their memory repertoire compared to the naïve repertoire of the same donor and show a shift in the usage of an important J gene responsible for the creation of long CDR3 (J6).
Theanti-HIVPGT121 broadly neutralizing antibodies have a 26 amino acid long heavy chain CDR3 with the original JH6 tyrosine repeat in the germline antibody and various mutations inside the CDR3 for 11 variants. We therefore study different B cell repertoires and the PGT antibodies as model for long CDR3.